Feature-based image registration approaches have historically fallen out of favour due to the difficulty of reliably extracting anatomical correspondences. Such challenges are now largely overcome thanks to recent advances in deep learning, with segmentation models capable of delivering reliable, fine-grained anatomical delineations in seconds. We leverage these advances to revive feature-based registration, creating alignment tools that are robust and anatomically grounded.

NimbleReg is a non-linear registration framework that operates on region boundary surfaces extracted from segmentations. Using a PointNet backbone, it is lightweight. Embedded in the stationary velocity field framework, it enables seamless multi-region blending to produce an overall diffeomorphic transformation.
Polaffini uses the centroids of segmented regions to efficiently estimate local affine matchings that are combined into an overall polyaffine transformation. Applicable both as standalone registration and as pre-alignment for non-linear methods, it is fast and robust, making it well-suited for integration into neuroimaging pipelines.
Diffusion MRI processing, from distortion correction to microstructure parameter estimation, is traditionally slow, taking hours per subject with iterative optimisation. Deep learning offers orders-of-magnitude speedups. We developed fast deep-learning methods addressing each stage of this pipeline, and explored protocol-agnostic approaches formicrostructure estimation.

This work introduces a protocol-agnostic method based on a PointNet architecture which is lightweight and possesses nice symmetries. Each voxel is represented as an unordered set of measurement tokens combining the signal with its acquisition parameters.
Eddeep is the first tool tackling eddy-current distortion correction with deep learning. It is composed of 2 models in sequence: 1) an image translator to restore correspondences between DW and b=0 volumes of very different contrast and, 2) a physics-based registration model to estimate the distortion and apply correction.

This work proposes a semi-supervised, physics-based deep-learning approach for fast susceptibility distortion correction of blip-up/blip-down acquisitions.
Population atlases capture the typical anatomy of a cohort, enabling spatial normalisation and population-level statistical analyses. Spatio-temporal atlases further model anatomical evolution over time. Their construction relies on non-linear registration and on the manipulation of geometric transformations under topology-preserving constraints.
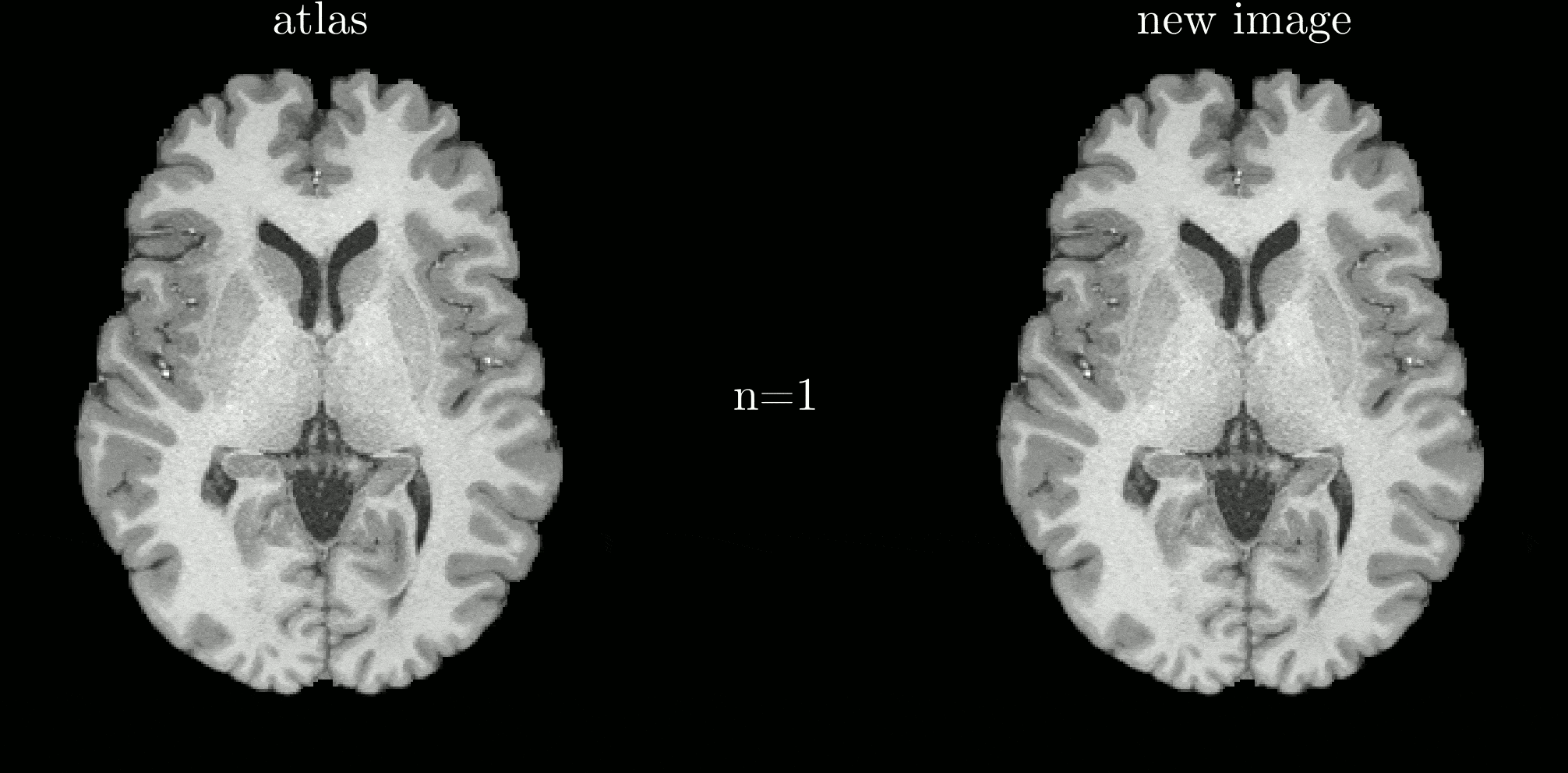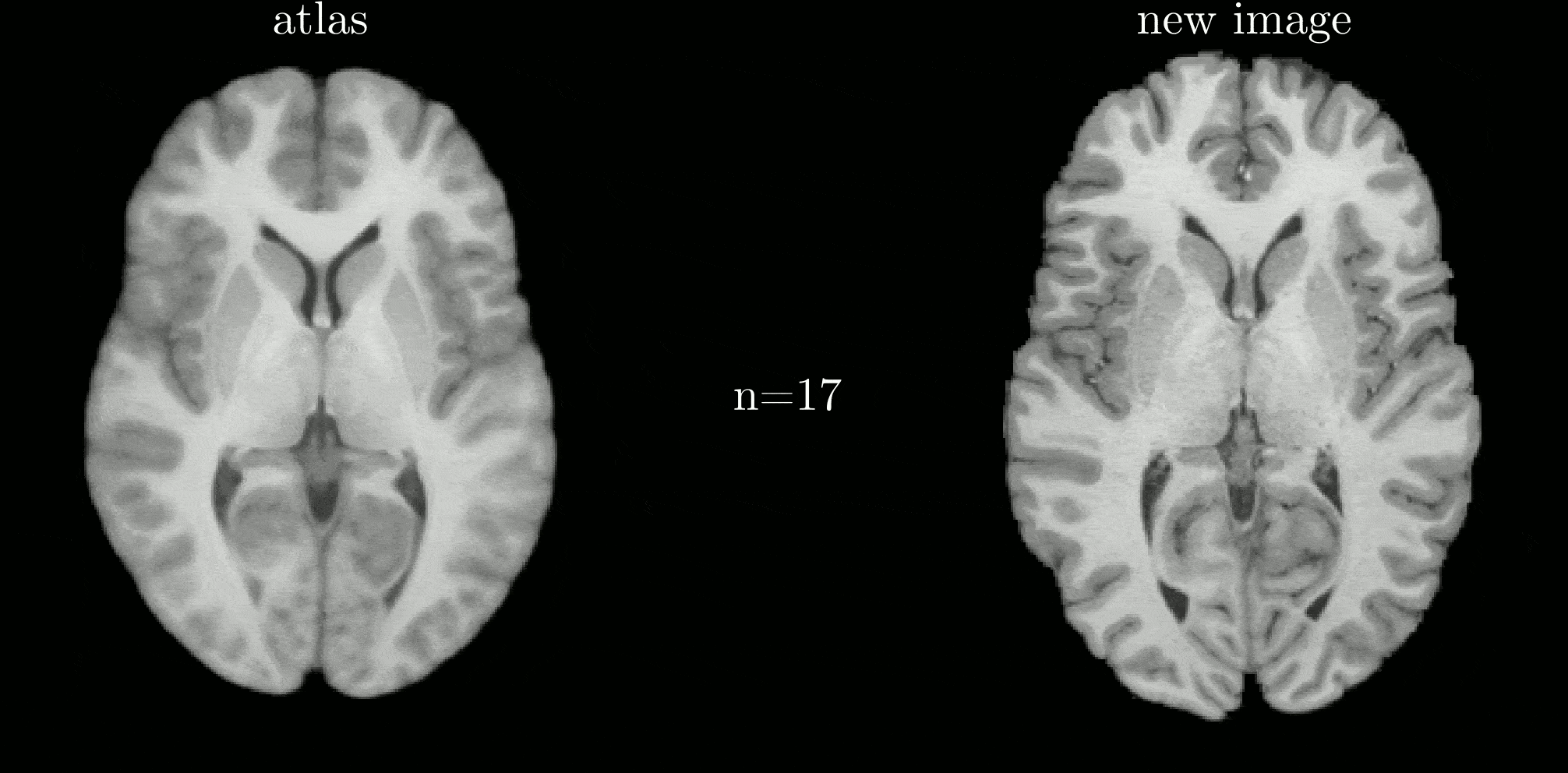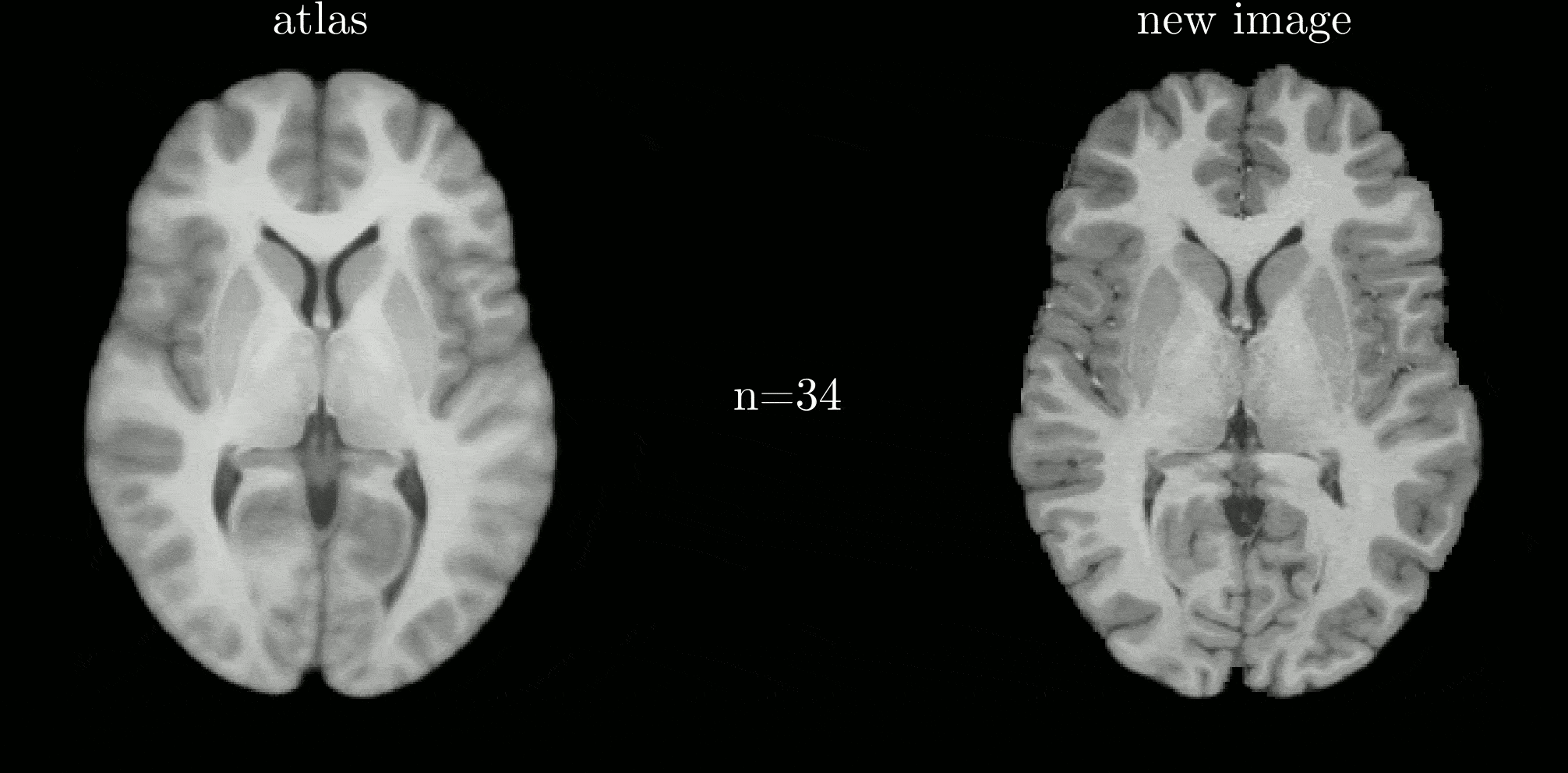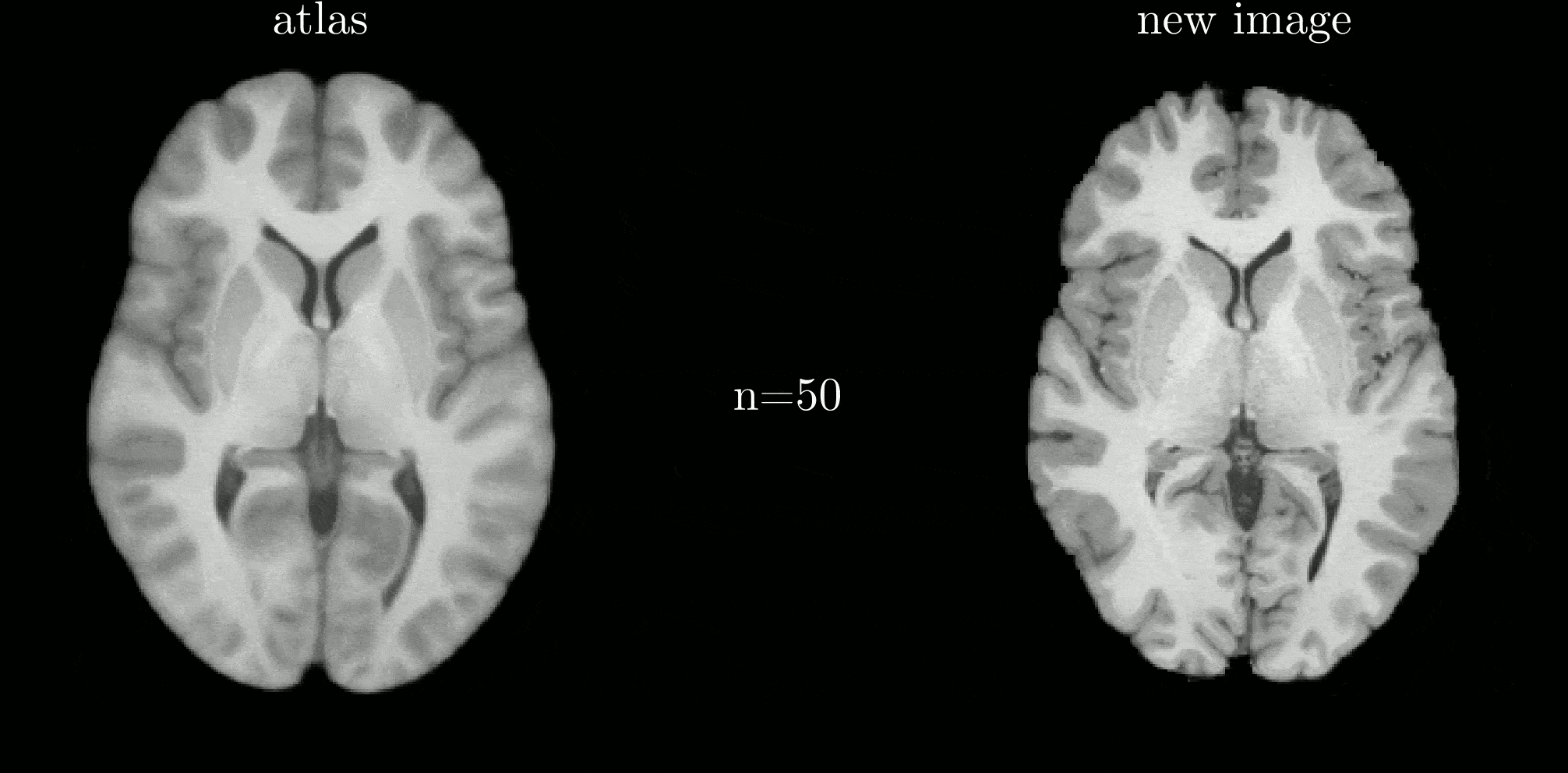
Classical atlas construction must be restarted from scratch when new data arrive. We introduce an iterative-centroid formulation that provides an online update rule to incorporate new subjects incrementally, based on principled manipulation of diffeomorphic transformations in the log-Euclidean framework.

Studying brain development requires atlases that simultaneously capture the population mean anatomy and its continuous evolution over time. This work proposes a diffeomorphic spatio-temporal atlasing method that jointly enforces geometric unbiasedness across individuals and temporal fidelity across developmental stages.